Adagio
From idea to running pipeline in minutes.
Build type-safe bioinformatics pipelines through a visual interface. Share pipelines as a single file. Run them anywhere. Every result maintains its entire history: inputs, parameters, and environment versions.
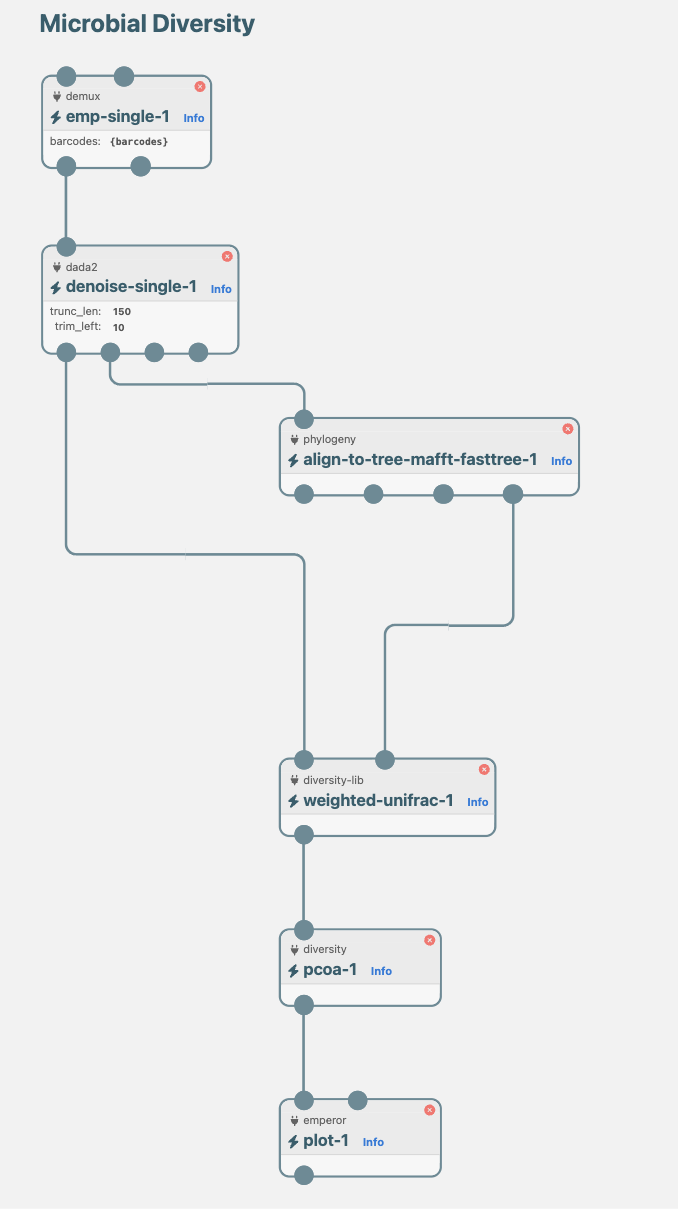
Build pipelines visually.
Pipelines are built through a click-and-drag interface, no code required. Bioinformatic actions are connected by edges defining data flow. The actions are all type safe and connections are only allowed to be made between matching data types. Data is validated based on types at run time ensuring that any pipeline you can build is valid and ready to be run. Pipelines can be saved and run locally, or started through the browser.
Browse community pipelines →Single-file. Run anywhere.
Share a pipeline the way you share a spreadsheet. Recipients need one command to run it: not a codebase, not a config directory, not a Slack thread of setup instructions.
pip install adagio-cli, get the .adg file,
run it. Parameters are exposed dynamically on the CLI. No browsing source
code to find what flags exist.
pip install adagio-cli wget \ -O "sample-metadata.tsv" \ "https://data.qiime2.org/2024.10/tutorials/moving-pictures/sample_metadata.tsv" wget https://docs.qiime2.org/2024.10/data/tutorials/moving-pictures/emp-single-end-sequences.qza adagio run adagio-playbook/denoise \ --input-seqs emp-single-end-sequences.qza \ --input-barcodes sample-metadata.tsv \ --param-barcodes barcode-sequence --param-trim-left 10 --param-trunc-len 140
execution: runtime: start: 2026-02-03T15:06:30.880957+00:00 end: 2026-02-03T15:07:51.538790+00:00 duration: 1 minute, 20 seconds, and 657833 microseconds execution_context: type: synchronous action: type: method plugin: 'environment:plugins:composition' action: ancombc2 inputs: - table: eb49efb6-ef8b-4ea6-acae-ab0bdc2e457e parameters: - metadata: 'metadata.tsv' - fixed_effects_formula: SampleType - random_effects_formula: null - reference_levels: - SampleType::Human Excrement Compost - p_adjust_method: holm - prevalence_cutoff: 0.1 - group: null - structural_zeros: false - asymptotic_cutoff: false - alpha: 0.05 - num_processes: 1 output-name: ancombc2_output environment: platform: Linux-6.8.0-1029-aws-x86_64-with-glibc2.39 python: |- 3.10.14 | packaged by conda-forge | (main, Mar 20 2024, 12:45:18) [GCC 12.3.0] framework: version: 2026.1.0 website: https://qiime2.org plugins: types: version: 2026.1.0 website: https://github.com/qiime2/q2-types composition: version: 2026.1.0+0.g4b3aa86.dirty website: https://github.com/qiime2/q2-composition python-packages: altair: 6.0.0 annotated-types: 0.7.0 anyio: 4.12.1 appdirs: 1.4.4 argcomplete: 3.6.3 ...
Your results remember everything.
Every result generated by Adagio is bundled with its complete history. Inputs, parameters, dependency versions, runtime, citations. The entire pipeline used to create any piece of data can be recreated with only that data.
Learn about provenance →Community Pipelines and Plugins
Over 400 bioinformatics tools and many pipelines are currently available for use with Adagio, with more being added every day. Use existing resources, create your own pipelines and plugins and publish them to the community, or save private workflows to your account.
Community →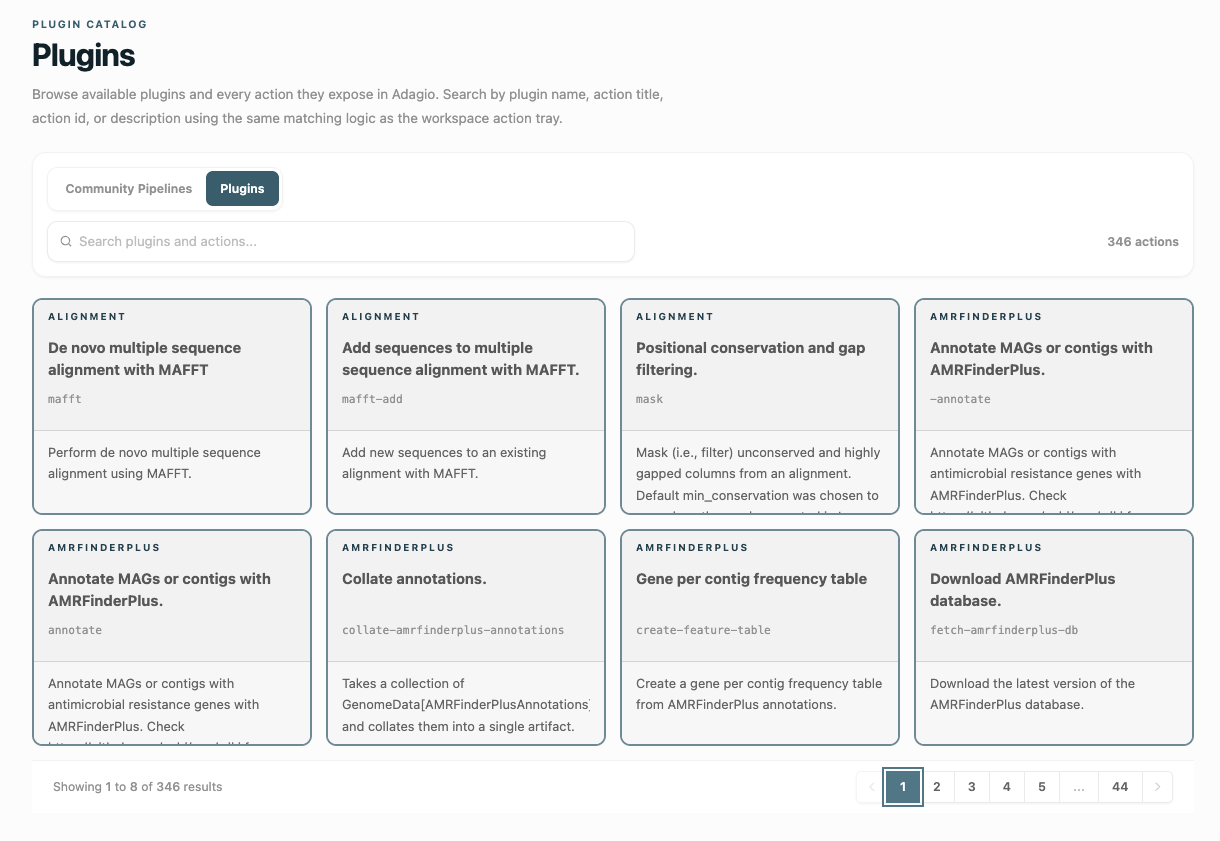
Join the waitlist to get early access.
By joining, you agree that Adagio may contact you about access and product updates. See our Privacy Policy.